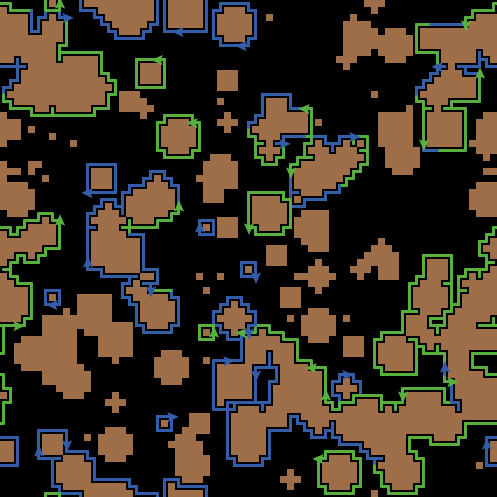

induction: proportion of mutant cells to be introduced (set to 0 for wild-type tissue simulations).immediate-stratification: true for enabling crowding release, induced by the double state cells (see main text).crowdingCutOff: number of neighbours above which the neighbourhood is considered "crowded".densityBias: set to true to enable cell density dependent feedbacks.delta: fate imbalance parameter of mutant cells (min:0, max:1) - p53delta and notchdelta in unified model.gamma: stratification rate (gammaWT, gammaMUT in upstream and unified models where mutant and WT cells have distinct stratification rates).lambda: division rate (lambdaWT, lambdaMUT in upstream and unified models where mutant and WT cells have distinct division rates).r: probability of symmetric division (rWT, rMUT in upstream and unified models where mutant and WT cells have distinct symmetric division probabilities).diffusion: set to true for allowing an extended neighbourhood.division-bias: set to true to constrain divisions along a single axis (for oesophageal simulations).sims-duration: simulation time in weeks.Set to a specific value for reproducibility predefinedSeed: set to zero for random seed.generate_world: set to true to generate and store grid state per week in csv format (Default: False).

generate_views: set to true to generate and store grid images per week (Default: True).The s file contains the following variables: To open it in NetLogo click on the Included Files menu ( SP_s should be opened first).
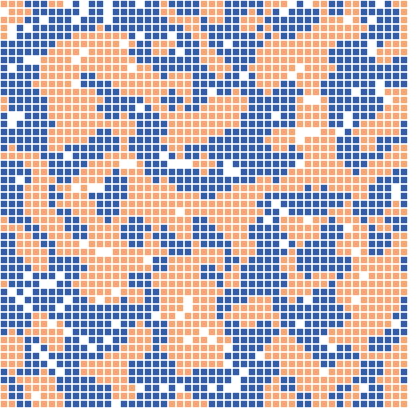

This may be opened using your prefered text editor or directly in the NetLogo environment. Model and simulation parameters can be set in the s file.
Netlogo northwestern how to#
$ java -Dfile.encoding=UTF-8 -cp netlogo-6.1.1.jar -model /path/to/unified/model/ogo -experiment unified -max-pxcor 99 -min-pxcor 0 -max-pycor 99 -min-pycor 0įor more information and examples on how to run NetLogo in headless mode see Configuration Parameters: $ cd path/to/netlogo/installation/directory (e.g.
Netlogo northwestern windows#
The following commands demonstrate how to run the SP spatial models in headless mode on Windows with NetLogo 6.1.1 installation, using a grid size of 100 x 100: Grid dimensions can be also set using the following arguments: The experiment setups for the SP spatial models have been already created and are saved in the model files. The netlogo-headless script expects the following required arguments:
Netlogo northwestern mac#
This is found in the root directory of your NetLogo installation and is named netlogo-headless.sh on Mac and Linux and netlogo-headless.bat on Windows. NetLogo offers command line usage by using the included command line script. Model parameters and simulation duration are defined in the s file (See configuration parameters) Option 2: Using the command line: Requirements
For a quick tutorial on NetLogo models please see On the interface tab, click on setup button to initialise grid, then click on the go "forever" button to run the model. Open NetLogo GUI and use the following steps:įile -> open -> select ogo file How to run: Option 1: Using NetLogo graphical interface Requirements analysis_output: Directories for storing analysis output plots and pickled variables.For enabling see generate_views, generate_ world parameters in s netlogo_output: Directories for storing output files - initially empty.s: helper functions for generating and saving output files.s: Configuration file for changing parameters to modify model behaviour.analysis: contains python scripts for analysing the simulation outputs.Įach model directory contains the following files:.unified: contains the model that enables both downstream and upstream feedback for mutant competition simulations.upstream: contains the upstream feedback model.downstream: contains the downstream feedback model.More detailed description of these models can be found in this manuscript by Kostiou et al. It also contains a collection of python scripts for analysing the outputs of model simulations. This repository contains the spatial non-neutral single progenitor models that simulate the collective behaviour of cells in epithelial tissues and the spread of mutant cells.
